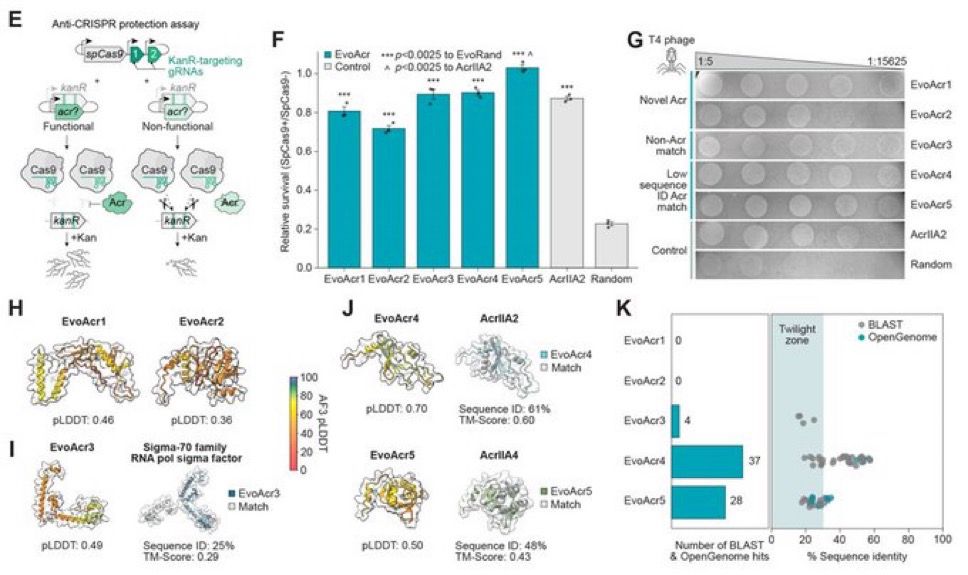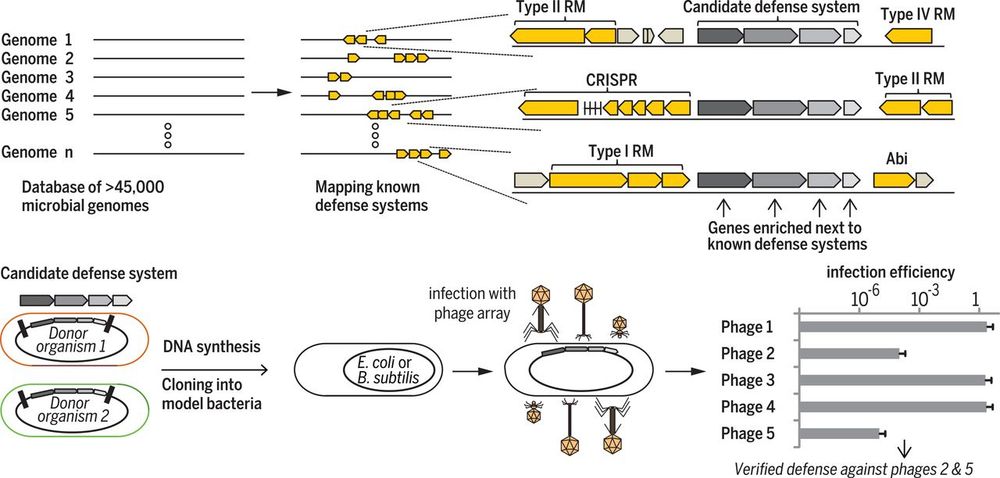

Today in @Nature, we share semantic design—a strategy for function-guided design with genomic language models that leverages genomic context to create de novo genes and systems with desired functions. 🧵
www.nature.com/articles/s41...
Today in @Nature, we share semantic design—a strategy for function-guided design with genomic language models that leverages genomic context to create de novo genes and systems with desired functions. 🧵
www.nature.com/articles/s41...