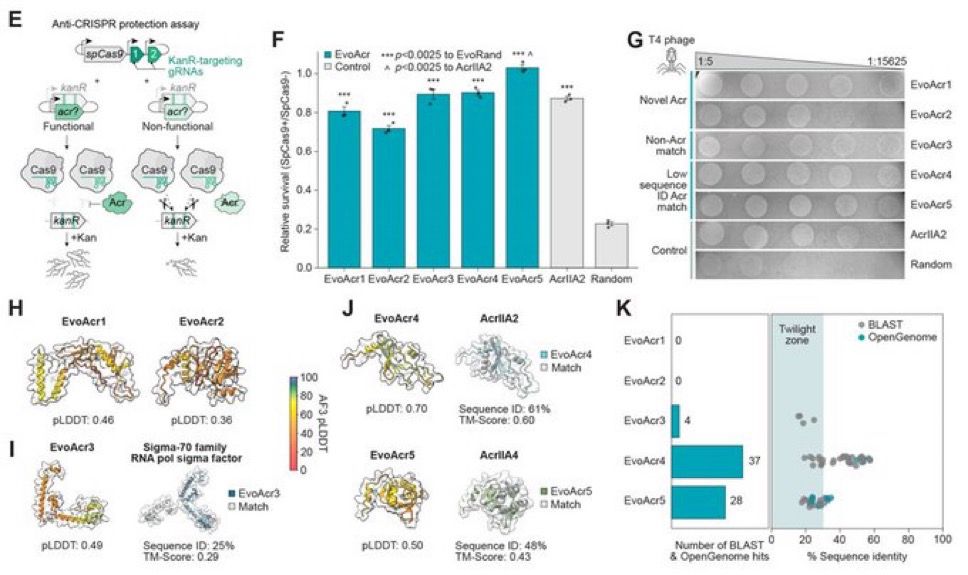
To learn more, check out the paper: www.nature.com/articles/s41...

To learn more, check out the paper: www.nature.com/articles/s41...

evodesign.org/syngenome/

evodesign.org/syngenome/





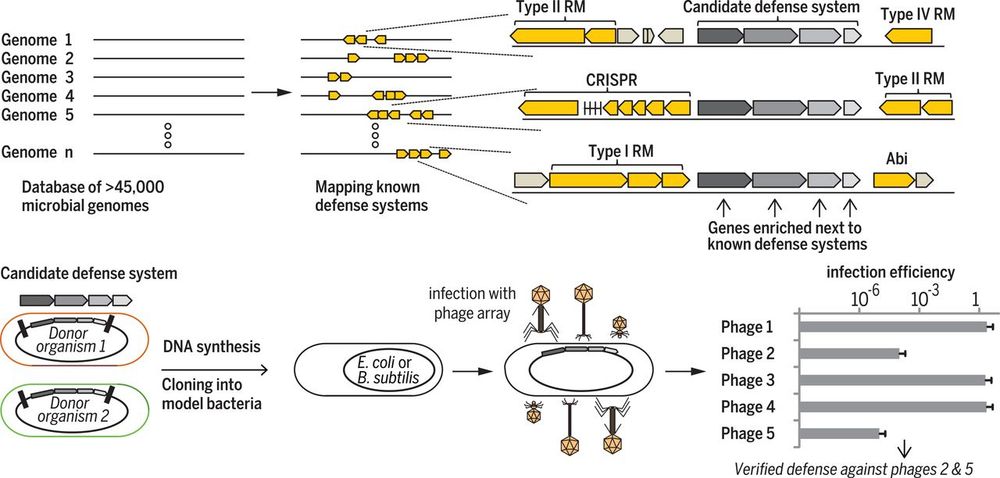

www.biorxiv.org/content/10.1...
evodesign.org/Semantic_Min...
www.biorxiv.org/content/10.1...
evodesign.org/Semantic_Min...