
http://www.type3secretionlab.es
#MicroSky #PlantScience #Pseudomonas
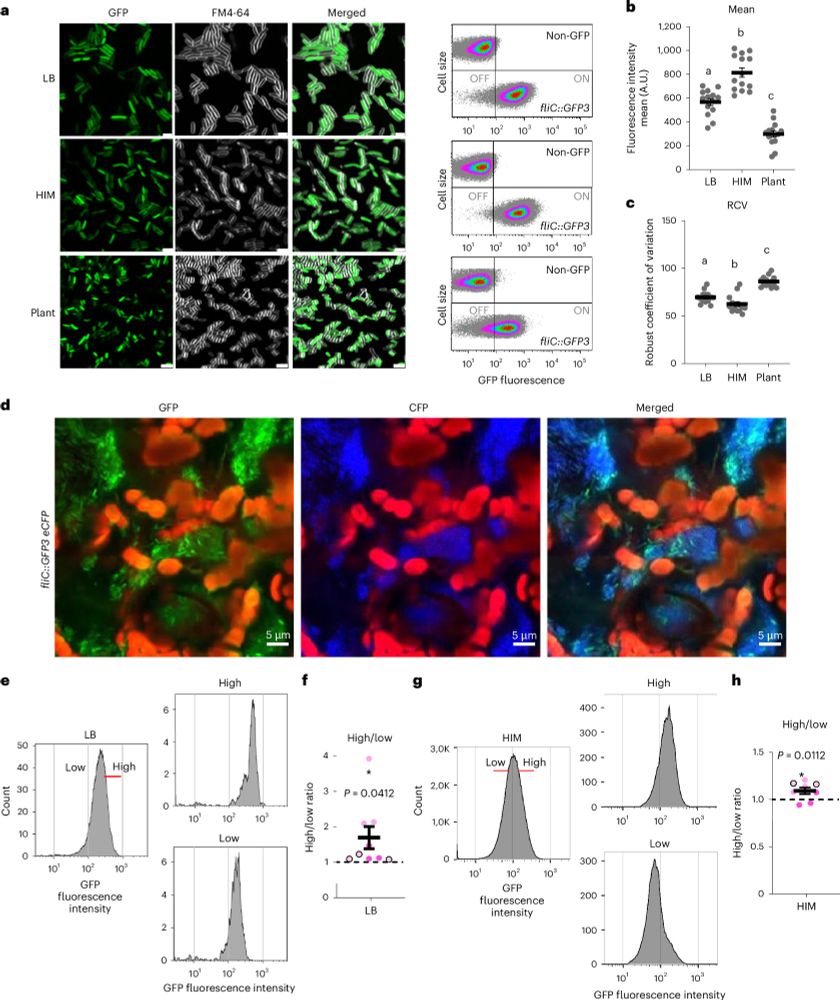
Our new research in Nature Microbiology uncovers the sophisticated teamwork of Pseudomonas syringae, a notorious plant pathogen.
🔗 rdcu.be/egczU
In this review, Elvira-González et al. describe how virus-induced small RNA synthesis and small RNA movement through plasmodesmata and phloem determine the outcome of viral infection in terms of disease and tolerance.
🔗 doi.org/10.1093/jxb/...
#PlantScience 🧪

In this review, Elvira-González et al. describe how virus-induced small RNA synthesis and small RNA movement through plasmodesmata and phloem determine the outcome of viral infection in terms of disease and tolerance.
🔗 doi.org/10.1093/jxb/...
#PlantScience 🧪
Salehimoghaddam et al.
nph.onlinelibrary.wiley.com/doi/10.1111/...
#plantscience

Salehimoghaddam et al.
nph.onlinelibrary.wiley.com/doi/10.1111/...
#plantscience
My group at @zmbp-tuebingen.bsky.social is offering a post-doctoral position (4 years). We look for a structural biologist with experience in Cryo-EM/Cryo-ET to investigate the mechanisms of host invasion by pathogenic fungi. Deadline February 28th!
uni-tuebingen.de/universitaet...

My group at @zmbp-tuebingen.bsky.social is offering a post-doctoral position (4 years). We look for a structural biologist with experience in Cryo-EM/Cryo-ET to investigate the mechanisms of host invasion by pathogenic fungi. Deadline February 28th!
uni-tuebingen.de/universitaet...
New paper out from the lab in Microbial Genomics, starting down the rabbit hole of IS elements in Paeudomonas syringae
www.microbiologyresearch.org/content/jour...

New paper out from the lab in Microbial Genomics, starting down the rabbit hole of IS elements in Paeudomonas syringae
www.microbiologyresearch.org/content/jour...
Don't miss it!
🕑 9:30 AM
📍 IHSM La Mayora Auditorium

Don't miss it!
🕑 9:30 AM
📍 IHSM La Mayora Auditorium
More information can be found here:
https://www.embo.org/funding/fellowships-grants-and-career-support/installation-grants/
#LifeSciences #research #funding

More information can be found here:
https://www.embo.org/funding/fellowships-grants-and-career-support/installation-grants/
#LifeSciences #research #funding

Abstract submission deadline: 1 May
Registration deadline: 31 May
https://meetings.embo.org/event/26-calcium-signaling
#EMBOCalciumSignaling #EMBOevents 🧪

Abstract submission deadline: 1 May
Registration deadline: 31 May
https://meetings.embo.org/event/26-calcium-signaling
#EMBOCalciumSignaling #EMBOevents 🧪

From CRISPR-triggered tRNA tail cleavage to temperature-responsive rRNA modifications, ribosome QC, and pathway-specific rRNA modification during biogenesis.
#RNAmods #Epitranscriptomics #Ribosome #Translation #tRNA #rRNA

From CRISPR-triggered tRNA tail cleavage to temperature-responsive rRNA modifications, ribosome QC, and pathway-specific rRNA modification during biogenesis.
#RNAmods #Epitranscriptomics #Ribosome #Translation #tRNA #rRNA
@dukasju.bsky.social @laurentterradot.bsky.social

@dukasju.bsky.social @laurentterradot.bsky.social


Baral and Brosché
nph.onlinelibrary.wiley.com/doi/10.1111/...

Baral and Brosché
nph.onlinelibrary.wiley.com/doi/10.1111/...
From @newphyt.bsky.social: shorturl.at/jYbOS
#PlantScience #PlantImmunity

From @newphyt.bsky.social: shorturl.at/jYbOS
#PlantScience #PlantImmunity

Our programme now features even more events, check it out ➡️ s.embl.org/2026-poster

Our programme now features even more events, check it out ➡️ s.embl.org/2026-poster

Full #openaccess
👉 doi.org/qktv
#PlantScience

Full #openaccess
👉 doi.org/qktv
#PlantScience

🇳🇵 Bishwash Thapa
🇮🇷 Zahra Nezamivand Chegini
🇨🇭 Sara Mitri (@saramitri.bsky.social)
🇺🇸 Lilian Caesar (@liliancaesar.bsky.social)
🇮🇹 Haseeb Manzoor
🇦🇹 Laurenz Holcik (@laurenz0908.bsky.social)
Free registration: cassyni.com/s/mvif-45
See you 👋

www.biorxiv.org/content/10.6...

www.biorxiv.org/content/10.6...
Discuss the latest approaches & discoveries in plant health with international keynote and local speakers.
APPLY by 30 March '26 ⬇️ Click link for more info
www.tsl.ac.uk/tsl-summer-c...

Discuss the latest approaches & discoveries in plant health with international keynote and local speakers.
APPLY by 30 March '26 ⬇️ Click link for more info
www.tsl.ac.uk/tsl-summer-c...


