https://www.cs.cit.tum.de/cmm
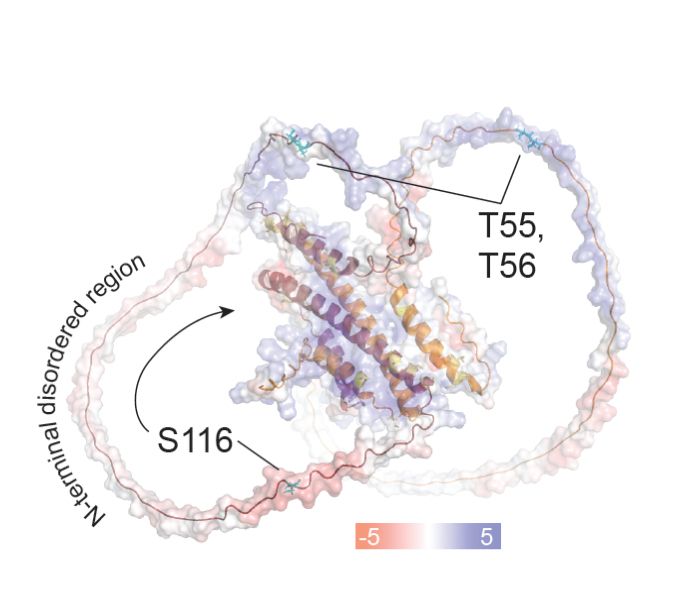


Existing tools often struggle with PTMs, yet PTMs are central to protein regulation and disease. We built a model for 19 amino acid-PTM combinations, leveraging data from in vivo experiments and from PROSPECT-PTM.

Existing tools often struggle with PTMs, yet PTMs are central to protein regulation and disease. We built a model for 19 amino acid-PTM combinations, leveraging data from in vivo experiments and from PROSPECT-PTM.
doi.org/10.1101/2025...

doi.org/10.1101/2025...



doi.org/10.1101/2024...

doi.org/10.1101/2024...






www.biorxiv.org/content/10.1...

www.biorxiv.org/content/10.1...

