
ORCID: 0000-0003-0994-5652
#CBIAS2025
#CBIAS2025
📄 arxiv.org/pdf/2507.01009 (1/N)
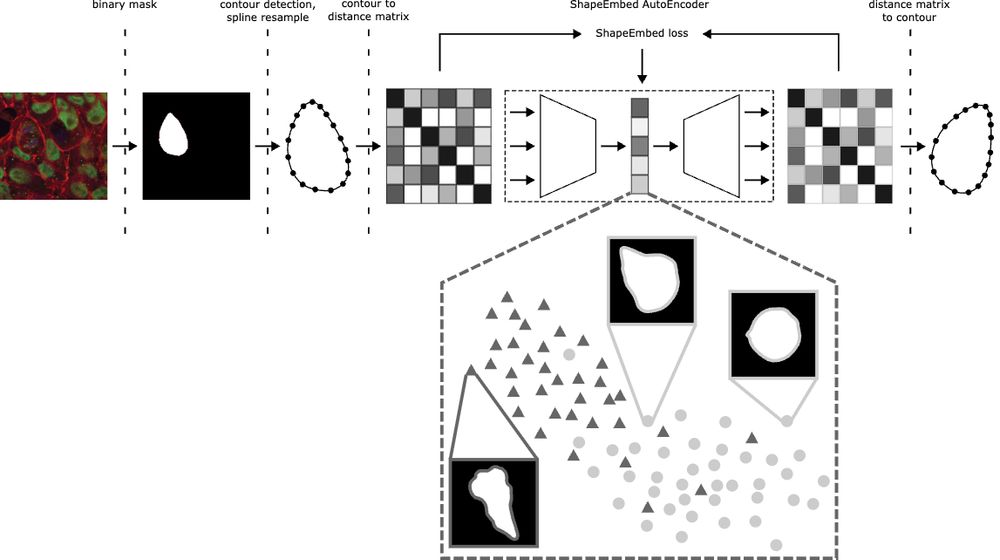
📄 arxiv.org/pdf/2507.01009 (1/N)
It's is not the end but the beginning of tons of papers to read, colleagues to contact, new collaborations to form. And most importantly, it's a 'see you soon' #CBIAS2026 🔬🥰

It's is not the end but the beginning of tons of papers to read, colleagues to contact, new collaborations to form. And most importantly, it's a 'see you soon' #CBIAS2026 🔬🥰

If you’d like to learn more about movement, the Python package I presented, my slides are live at neuroinformatics.dev/slides-movem...
If you’d like to learn more about movement, the Python package I presented, my slides are live at neuroinformatics.dev/slides-movem...







#CBIAS2025


#CBIAS2025








Loads of new features, most excitingly to me deep learning segmentation algorithms including Omnipose (for cells), and Spotiflow (for spots) #bioimaging

Loads of new features, most excitingly to me deep learning segmentation algorithms including Omnipose (for cells), and Spotiflow (for spots) #bioimaging

Tracking is hard because segmentation is hard, especially in whole zebra fish embryos: Ultrack can help github.com/royerlab/ult...


Tracking is hard because segmentation is hard, especially in whole zebra fish embryos: Ultrack can help github.com/royerlab/ult...





