Identifies diff. expressed ion channels/networks via transcriptomics, understanding tumor behavior.

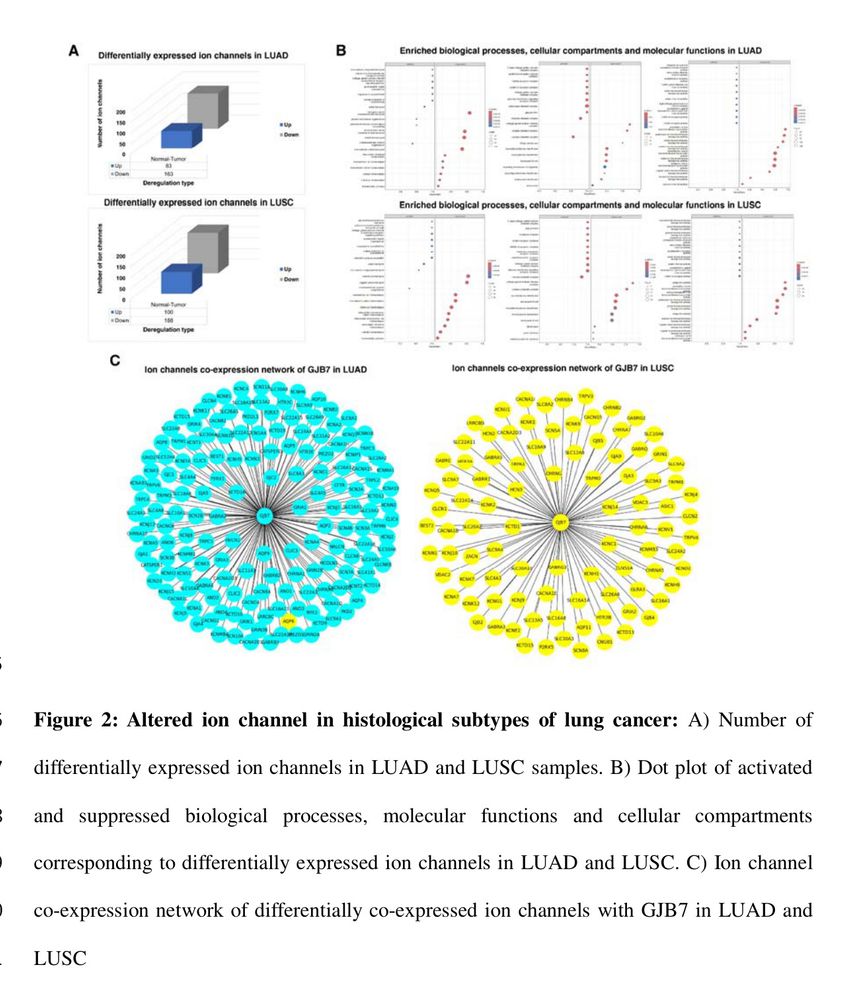


Identifies diff. expressed ion channels/networks via transcriptomics, understanding tumor behavior.
PatchDNA: Flexible, vocab-free DNA patching via evolution guides better LMs.




PatchDNA: Flexible, vocab-free DNA patching via evolution guides better LMs.
Examines diffusion models to relate learned functions to thermodynamics for protein folding, testing the ability to capture learned energies.




Examines diffusion models to relate learned functions to thermodynamics for protein folding, testing the ability to capture learned energies.








Generates short antimicrobial peptides using a classifier-guided diffusion model, optimizing for antimicrobial activity and low toxicity.




Generates short antimicrobial peptides using a classifier-guided diffusion model, optimizing for antimicrobial activity and low toxicity.
Single-cell & spatial omics via disentangled learning; cross-modal prediction & spatial mapping.




Single-cell & spatial omics via disentangled learning; cross-modal prediction & spatial mapping.
Denoises fluoro-microscopy via implicit nets & neighbor-sample; preserves detail @ low SNR.




Denoises fluoro-microscopy via implicit nets & neighbor-sample; preserves detail @ low SNR.
DNA methylation models: Accuracy/efficiency trade-offs in 3 CpG methylation dynamics methods.




DNA methylation models: Accuracy/efficiency trade-offs in 3 CpG methylation dynamics methods.
Ecosystem consolidates protein design models, workflows, data, and visualization into a scalable, open-source platform for experts and non-experts.




Ecosystem consolidates protein design models, workflows, data, and visualization into a scalable, open-source platform for experts and non-experts.
Cell-type specific spatial ligand-receptor analysis reveals signaling mechanisms.




Cell-type specific spatial ligand-receptor analysis reveals signaling mechanisms.
NanoporeDB: Structural database of multimeric protein nanopore models for functional interpretation and application advancement.


NanoporeDB: Structural database of multimeric protein nanopore models for functional interpretation and application advancement.
Deep learning found bacterial UPOs, bypassing eukaryotic issues & boosting use.




Deep learning found bacterial UPOs, bypassing eukaryotic issues & boosting use.




Leverages language models to create a new 20-letter protein alphabet. This enables fast, sensitive homology searches without structural data.




Leverages language models to create a new 20-letter protein alphabet. This enables fast, sensitive homology searches without structural data.
Fine-tuning ProtLMs aids TF ID & DNA binding. Attn weights/motifs boost interpretability.




Fine-tuning ProtLMs aids TF ID & DNA binding. Attn weights/motifs boost interpretability.
NMR cells show increased RNA turnover, particularly in aging-related pathways, suggesting RNA degradation modulates gene expression during aging.




NMR cells show increased RNA turnover, particularly in aging-related pathways, suggesting RNA degradation modulates gene expression during aging.
Gathers/unifies compound-protein data, filters, harmonizes IDs, computes binding values.


Gathers/unifies compound-protein data, filters, harmonizes IDs, computes binding values.
Multivariate & functional regression for fine-mapping traits across cells.




Multivariate & functional regression for fine-mapping traits across cells.
Antibiotic stress & bacterial survival: Morphology links survival under 168 antibiotics; ID'ing action modes via neural net.




Antibiotic stress & bacterial survival: Morphology links survival under 168 antibiotics; ID'ing action modes via neural net.
FGT spatial atlas: mol. features linked to TME. IDs HPV/TGFβ, oncogene phenotypes, & novel PCD niche for prognosis.




FGT spatial atlas: mol. features linked to TME. IDs HPV/TGFβ, oncogene phenotypes, & novel PCD niche for prognosis.
LLMs interpret rare disease variants via nat. lang. & autonomous genomic DB workflows.




LLMs interpret rare disease variants via nat. lang. & autonomous genomic DB workflows.
Infers gene networks by creating a prior interaction graph using text embeddings optimized with a graph neural network and scRNA-seq data.



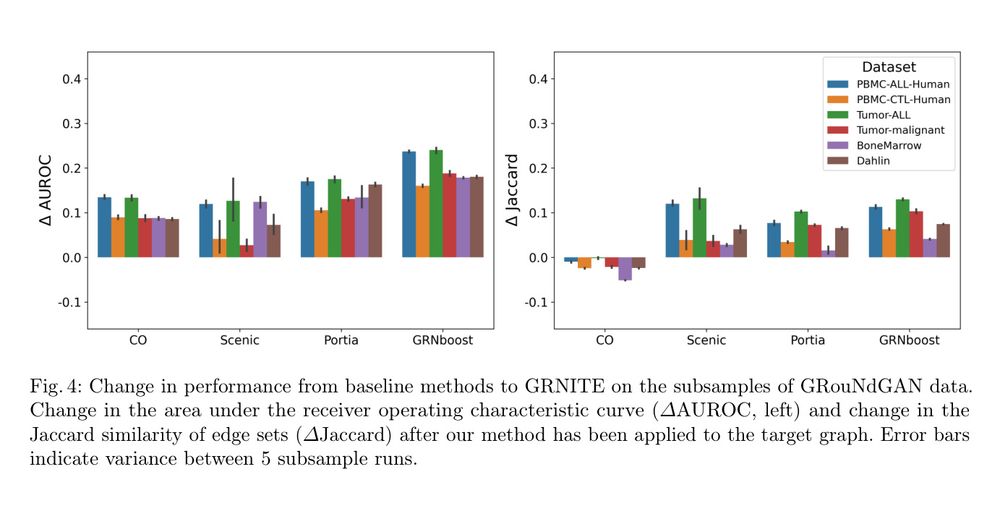
Infers gene networks by creating a prior interaction graph using text embeddings optimized with a graph neural network and scRNA-seq data.
Combines dimension reduction & clustering via Gaussian latent variable models, applicable to genomics data like gene expression and scRNA-seq.




Combines dimension reduction & clustering via Gaussian latent variable models, applicable to genomics data like gene expression and scRNA-seq.





