suragnair.github.io

careers.gene.com/us/en/job/20...

careers.gene.com/us/en/job/20...

You can find the preprint here: www.biorxiv.org/content/10.1...
12/

You can find the preprint here: www.biorxiv.org/content/10.1...
12/
Nona’s flexible masking schemes open new directions, and there’s much more to explore. 11/
Nona’s flexible masking schemes open new directions, and there’s much more to explore. 11/


They allow parallel decoding, with competitive performance at fewer generation steps. 8/


They allow parallel decoding, with competitive performance at fewer generation steps. 8/







Nona operates on both DNA sequence and functional genomics tracks. Task-specific masking configurations recover familiar model types, and its flexibility enables entirely new approaches! 2/
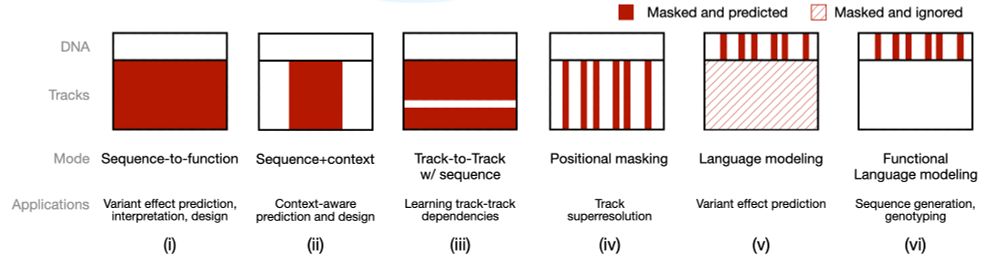
Nona operates on both DNA sequence and functional genomics tracks. Task-specific masking configurations recover familiar model types, and its flexibility enables entirely new approaches! 2/

