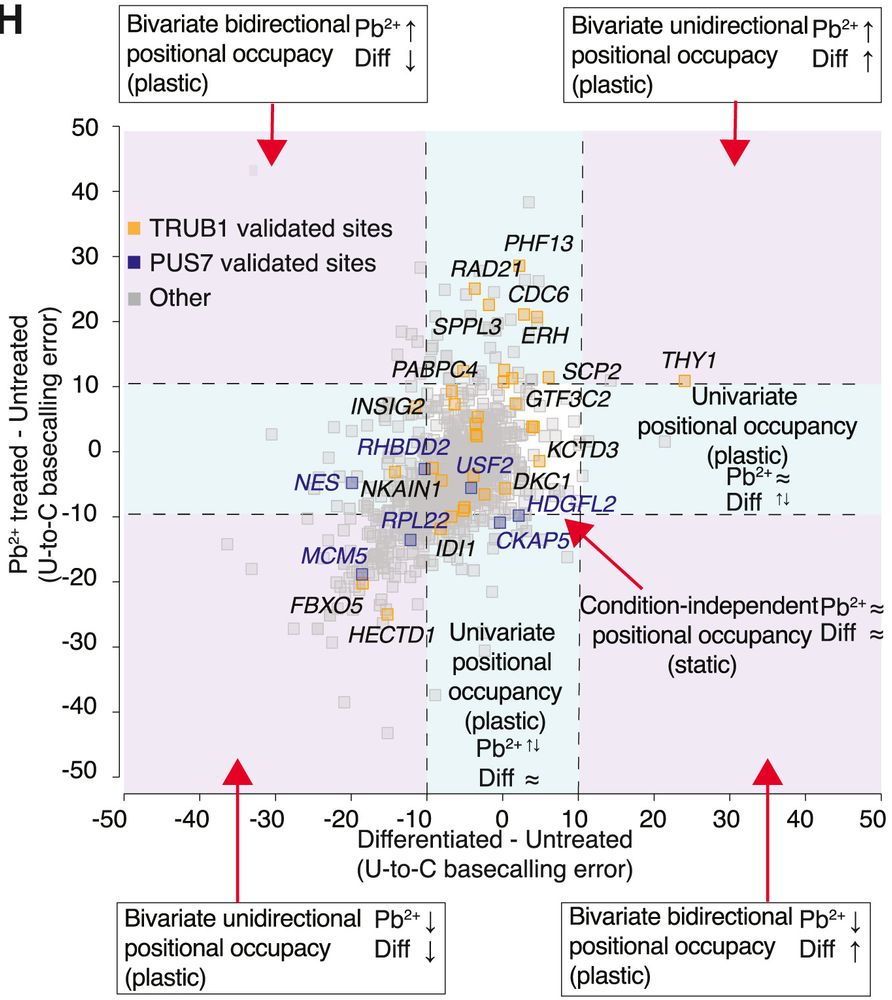
☑️ Jurkat-specific ψ-sites hit oncogenic and immune activation genes
☑️ Primary T cell–specific ψ-sites appear on trafficking and calcium signaling genes
🧬 Immortalized cells also showed more clustered ψ patterns, hinting at altered control compared to primary T cells.

☑️ Jurkat-specific ψ-sites hit oncogenic and immune activation genes
☑️ Primary T cell–specific ψ-sites appear on trafficking and calcium signaling genes
🧬 Immortalized cells also showed more clustered ψ patterns, hinting at altered control compared to primary T cells.
Most differences came from gene expression, not ψ-site selection.
This process seems to be tightly conserved.

Most differences came from gene expression, not ψ-site selection.
This process seems to be tightly conserved.
🚨 New paper published in RNA!
Scientists often say anecdotally that RNA modifications are disrupted in immortalized cells — but no one’s really tested it.
So we did: ψ-mapping in primary T cells vs. Jurkat cells using direct RNA-seq.
📄 tinyurl.com/TCellPsiRNA
#nanopore #RNA #pseudouridine
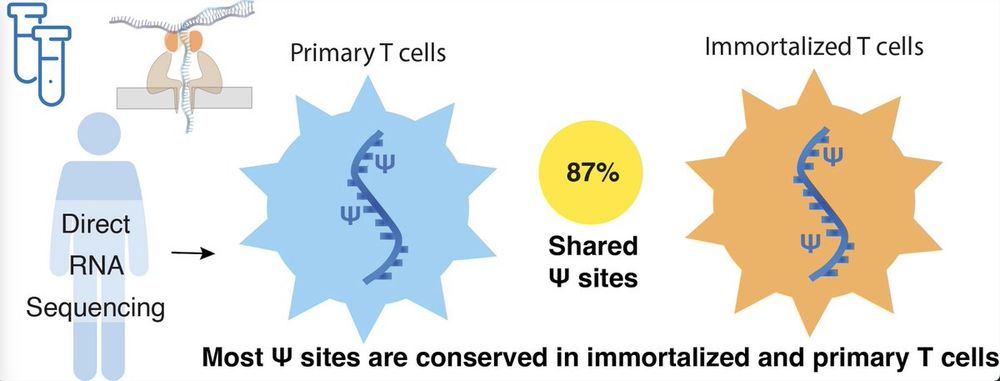
🚨 New paper published in RNA!
Scientists often say anecdotally that RNA modifications are disrupted in immortalized cells — but no one’s really tested it.
So we did: ψ-mapping in primary T cells vs. Jurkat cells using direct RNA-seq.
📄 tinyurl.com/TCellPsiRNA
#nanopore #RNA #pseudouridine