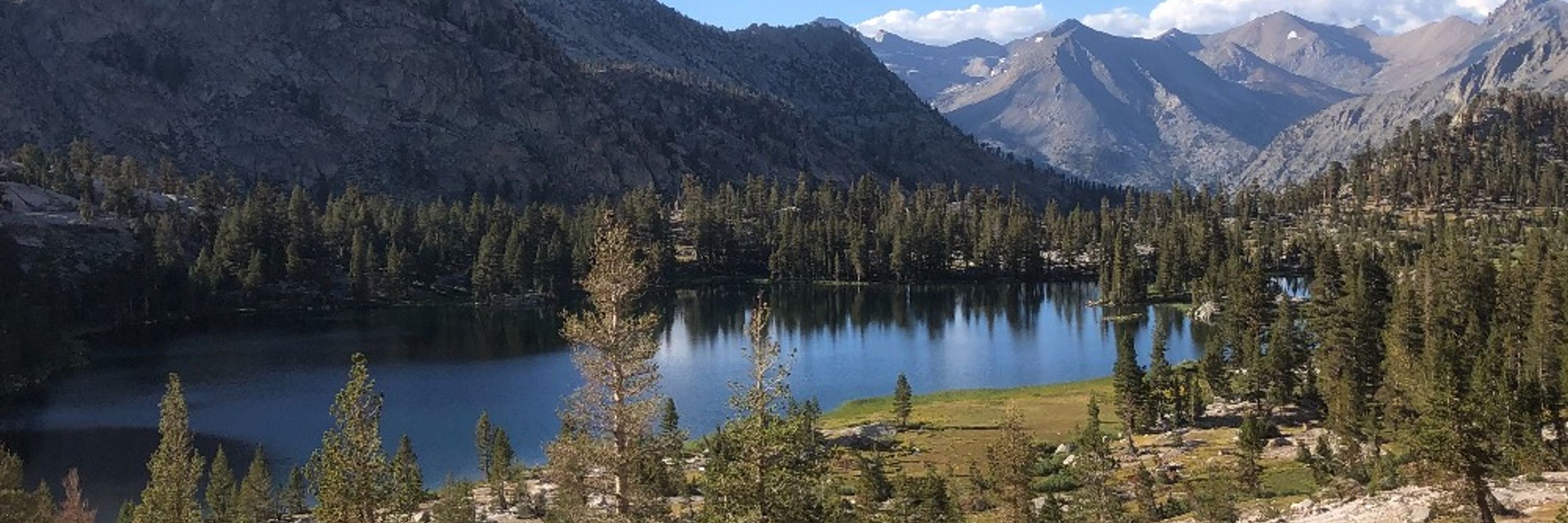

% of microbiome (& changes) aren't identifiable from HTS, but stacked barplots are helpful for generating clues esp if big shifts
data snooping has implications for error rate control. there isn't a hypothesis you're interested in?
good luck & enjoy xx
% of microbiome (& changes) aren't identifiable from HTS, but stacked barplots are helpful for generating clues esp if big shifts
data snooping has implications for error rate control. there isn't a hypothesis you're interested in?
good luck & enjoy xx
1. away from obsessing over alpha and beta diversity comparisons
2. towards comparisons that are less sensitive to rare/undetected species (than diversity)
So 📈
🥳
1. away from obsessing over alpha and beta diversity comparisons
2. towards comparisons that are less sensitive to rare/undetected species (than diversity)
So 📈
🥳
(Sorry -- I'm at a workshop today or I'd do it myself)
(Sorry -- I'm at a workshop today or I'd do it myself)
Feel free to open a feature request. We'll see what we can do.
github.com/statdivlab/r...
Feel free to open a feature request. We'll see what we can do.
github.com/statdivlab/r...
How do I describe data with a lot of variance? High-variance.
How do I describe data where the totals convey complex information about an unknown quantity I care about? (abundance)
I don't. I just state my assumptions.
How do I describe data with a lot of variance? High-variance.
How do I describe data where the totals convey complex information about an unknown quantity I care about? (abundance)
I don't. I just state my assumptions.
😽😽 7/6
😽😽 7/6
1. choose something meaningful to estimate
2. choose a sensible way to estimate it
3. choose tests that control Type 1 error
That's what we will keep doing, even if anonymous reviewers insist on buzzwords.
6/6
1. choose something meaningful to estimate
2. choose a sensible way to estimate it
3. choose tests that control Type 1 error
That's what we will keep doing, even if anonymous reviewers insist on buzzwords.
6/6
2/6
2/6

