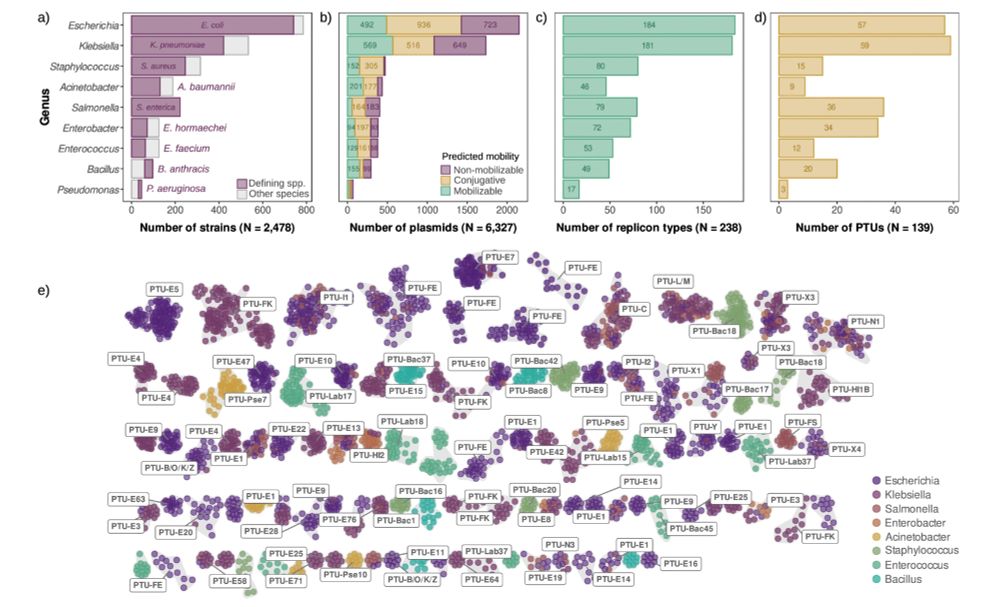
Bacterial and plasmid evolution 🫧🧬💻

We analyzed 1,598 plasmids (NCBI) 💻 to detect mutations arising during culture growth by comparing sequencing reads to consensus assemblies.
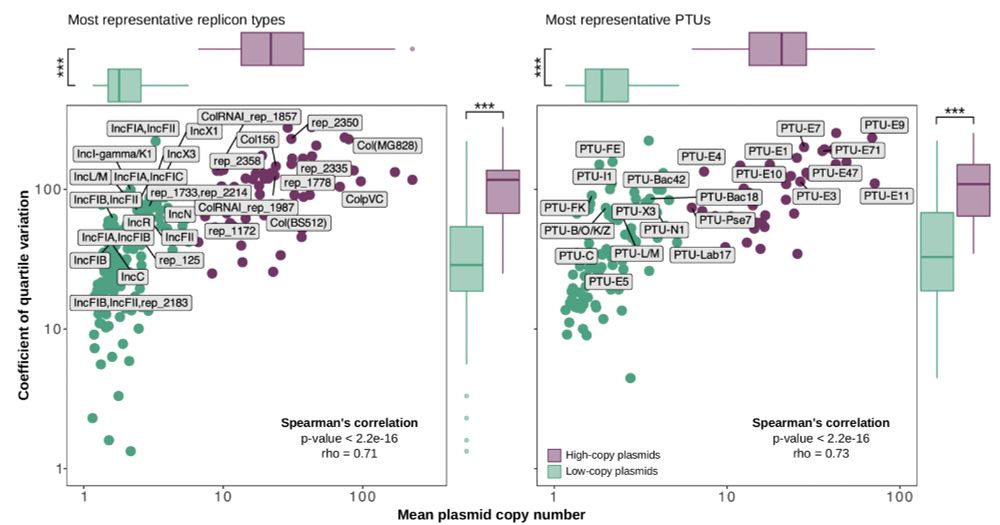
High-copy plasmids exhibited significantly higher mutation rates, just like theory (and experiments!) predicted!
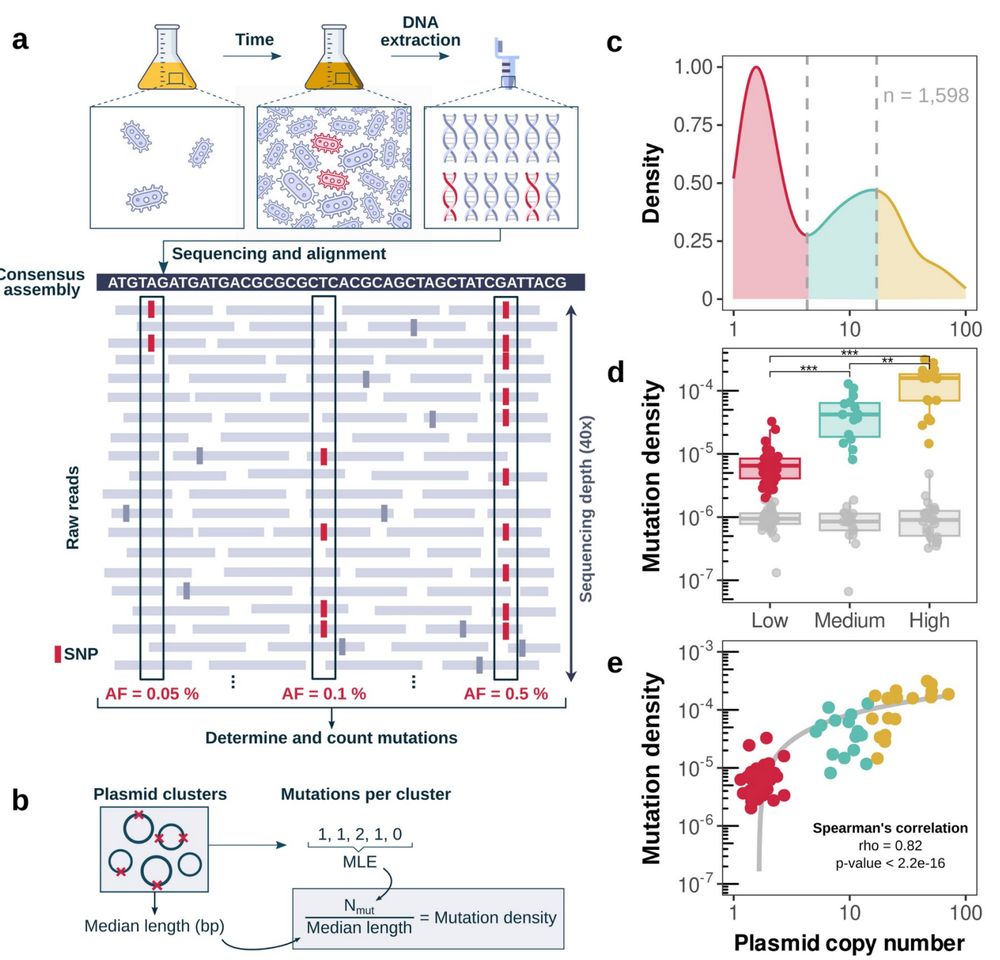
We analyzed 1,598 plasmids (NCBI) 💻 to detect mutations arising during culture growth by comparing sequencing reads to consensus assemblies.
High-copy plasmids exhibited significantly higher mutation rates, just like theory (and experiments!) predicted!
At more copies, mutation supply dominates drift! 💥
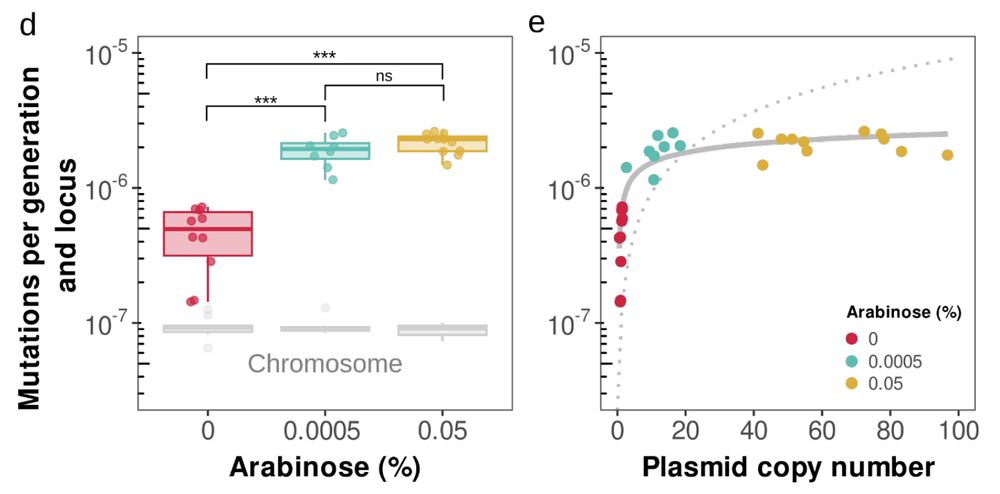
At more copies, mutation supply dominates drift! 💥
We established three PCN conditions: low (~1), medium (~10), and high (~60 copies). Lines were evolved for 30 days under severe bottlenecks.
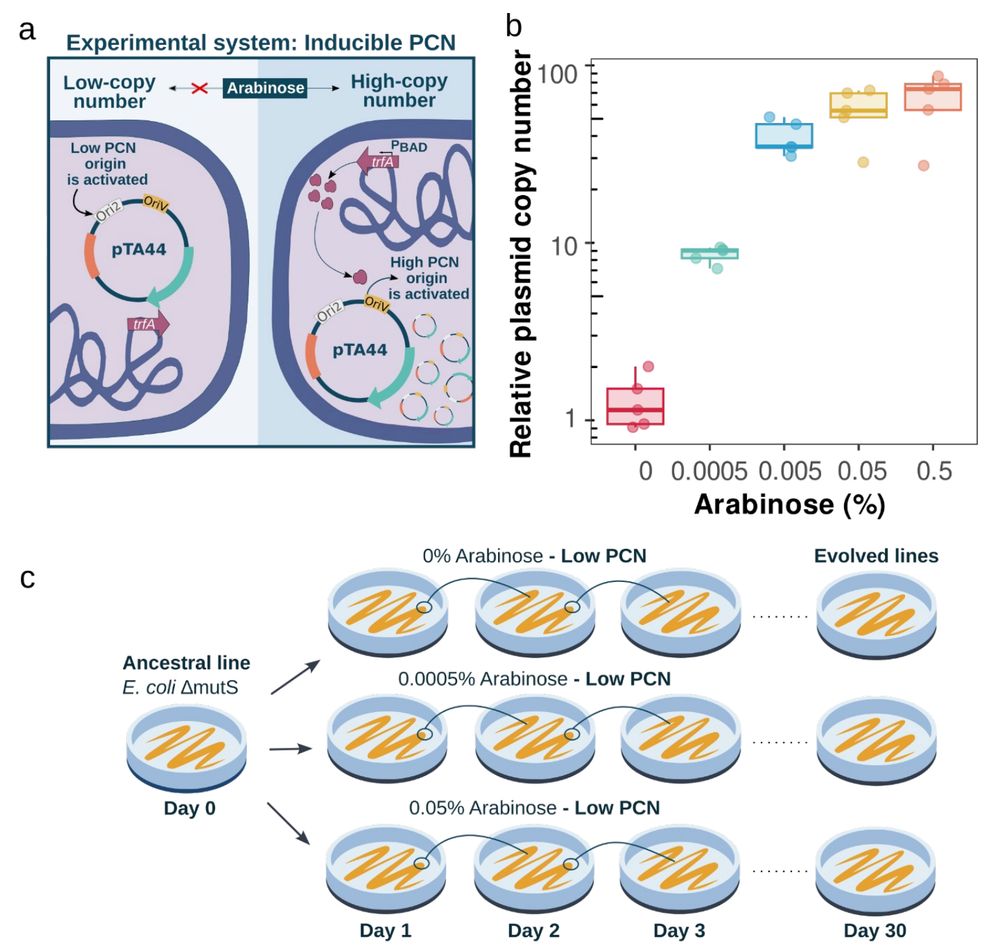
We established three PCN conditions: low (~1), medium (~10), and high (~60 copies). Lines were evolved for 30 days under severe bottlenecks.
Prediction: the mutation rate rises with PCN, but with diminishing returns, increasing logarithmically.
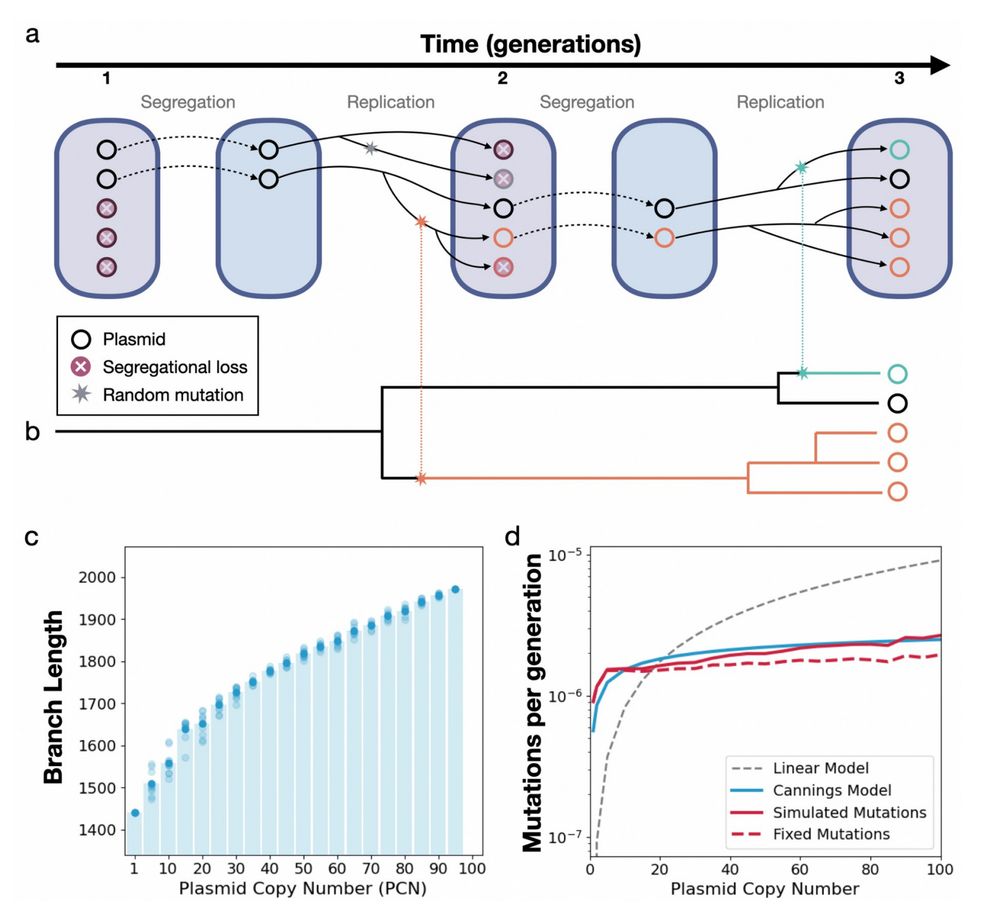
Prediction: the mutation rate rises with PCN, but with diminishing returns, increasing logarithmically.
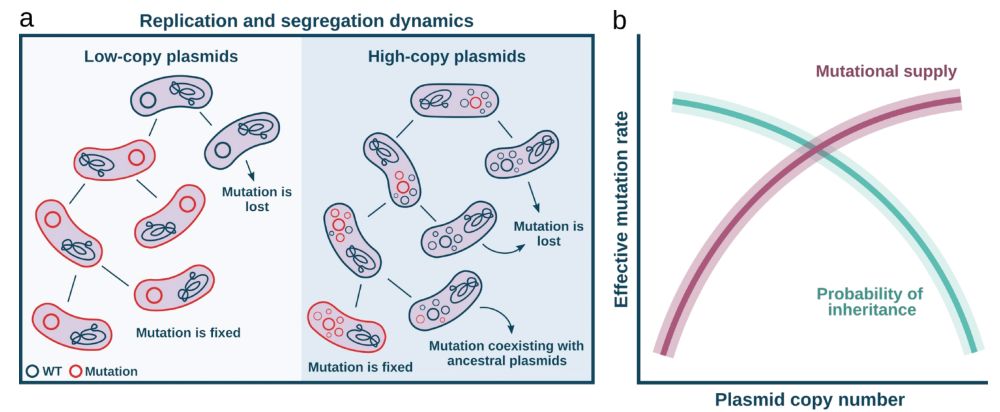
🔹 More copies = more chances to mutate 📈
🔹 But random segregation = drift, purging mutations 📉
So mutation supply ↑ with PCN, but the probability of fixation ↓
Which force dominates: the boost in mutation rate or the loss from segregational drift?
🔹 More copies = more chances to mutate 📈
🔹 But random segregation = drift, purging mutations 📉
So mutation supply ↑ with PCN, but the probability of fixation ↓
Which force dominates: the boost in mutation rate or the loss from segregational drift?