

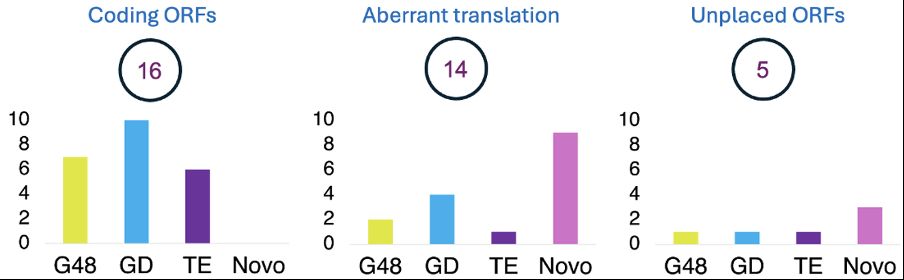








APPRIS does get it right, but only in RefSeq because RefSeq annotates the downstream ATG (169 aa isoform):

APPRIS does get it right, but only in RefSeq because RefSeq annotates the downstream ATG (169 aa isoform):
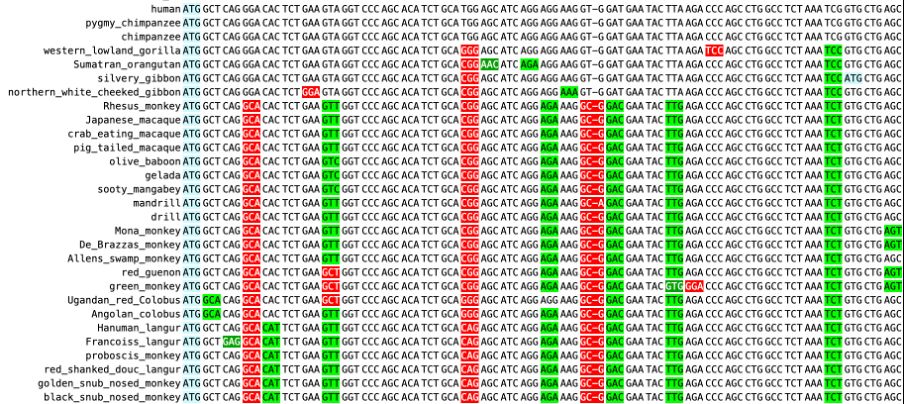
ADH6 seems to be an old world monkey duplication of ADH7, which has mutated a LOT.

ADH6 seems to be an old world monkey duplication of ADH7, which has mutated a LOT.
And that winds up our work in @gencodegenes.bsky.social
It has been fun.
Work carried out by Miguel Maquedano and @danielcerdan.bsky.social

And that winds up our work in @gencodegenes.bsky.social
It has been fun.
Work carried out by Miguel Maquedano and @danielcerdan.bsky.social
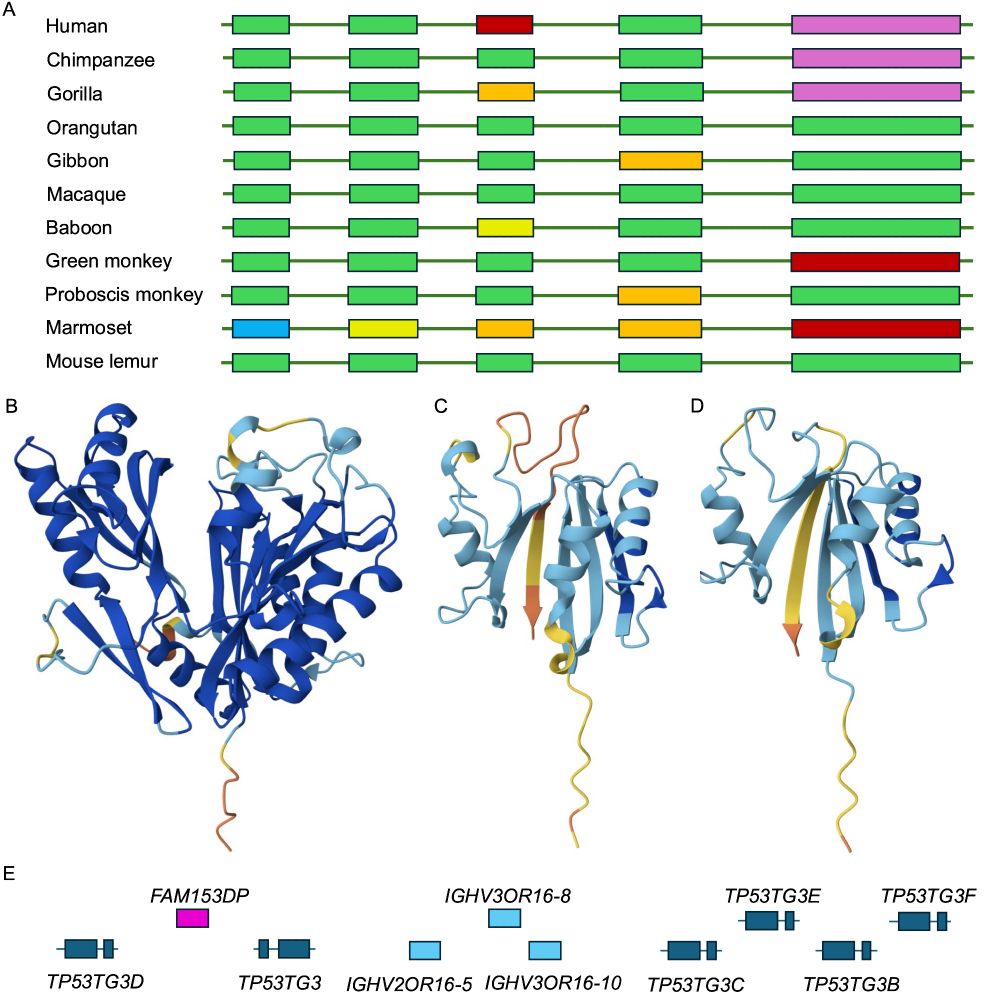
With the exception of the agreements between UniProtKB and Ensembl/GENCODE (mostly Ig/TcR fragments), most genes outside of the 3-way intersection were tagged as potential non-coding.
These are probably not coding genes.

With the exception of the agreements between UniProtKB and Ensembl/GENCODE (mostly Ig/TcR fragments), most genes outside of the 3-way intersection were tagged as potential non-coding.
These are probably not coding genes.
With the caveat that the Ensembl/GENCODE set is also the worst reference set for readthrough "coding" genes (see start of thread).

With the caveat that the Ensembl/GENCODE set is also the worst reference set for readthrough "coding" genes (see start of thread).

We merged and compared @ensembl.org / @gencodegenes.bsky.social , RefSeq and UniProtKB coding genes and investigated the agreements and discrepancies.
Details of what we found in the thread ...

We merged and compared @ensembl.org / @gencodegenes.bsky.social , RefSeq and UniProtKB coding genes and investigated the agreements and discrepancies.
Details of what we found in the thread ...

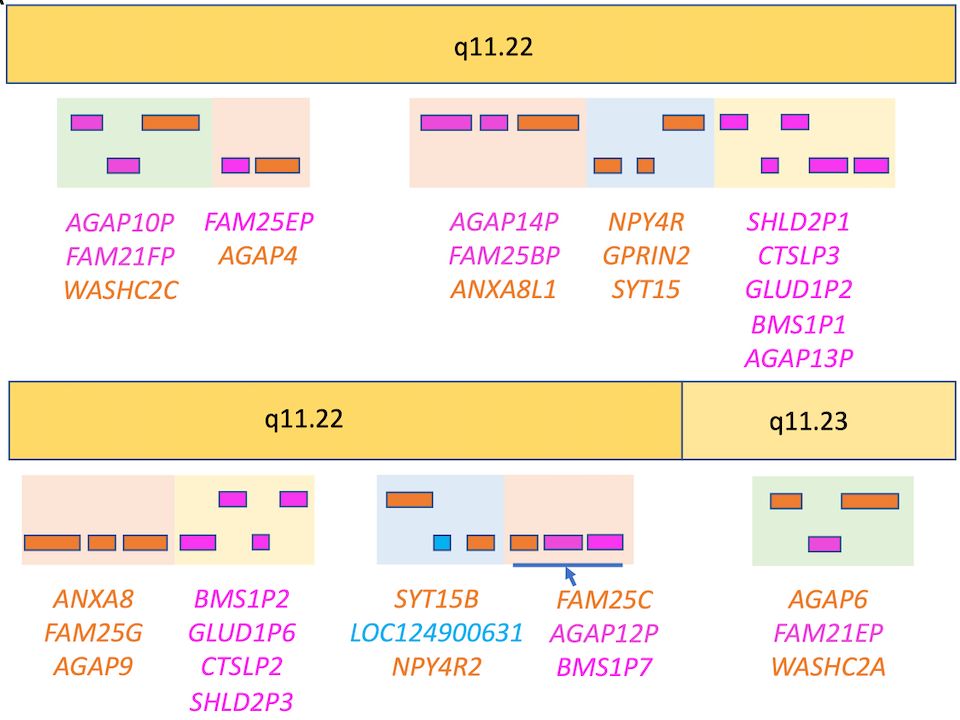
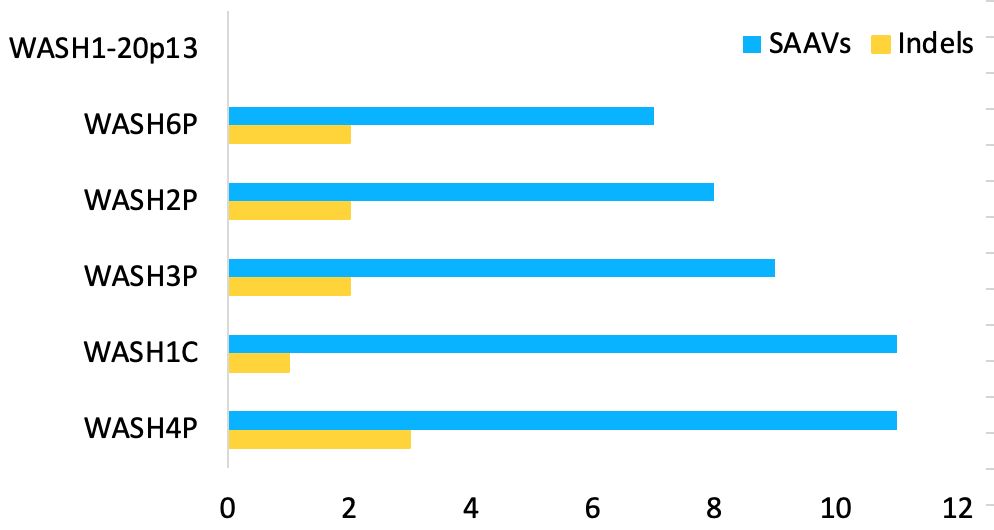
All 12 WASH12C paralogues are pseudogenes.

All 12 WASH12C paralogues are pseudogenes.
@bioinfoadv.bsky.social

@bioinfoadv.bsky.social
![The percentage of the genes in different gene sets that are tagged as potential non-coding genes. The genes in each set are those shown in figure 1B, “GENCODE [G]” are all Ensembl/GENCODE genes, “RefSeq [R]” are all RefSeq genes, and “UniProtKB [U]” are all UniProtKB genes. “G. not intersect” are all Ensembl/GENCODE genes that are not in the intersection between the three sets. “R. not intersect” are all RefSeq genes that are not in the intersection between the three sets. “U. not intersect” are all UniProtKB genes that are not in the intersection between the three sets.](https://cdn.bsky.app/img/feed_thumbnail/plain/did:plc:ea24pi345ceviookqvbeicoc/bafkreifxbp4dxujsvaanepm3rue4uf7fxjnkb4dbgchc654rplqoiphgyy@jpeg)


