Trying to master Dad mode 👨🍼
See what I'm up to here: https://github.com/mbhall88
If you’ve used nohuman before, I highly recommend updating to v0.5.0 and re-downloading the new DB.
Repo: github.com/mbhall88/nohuman
Keep your metagenomes clean 🧹🧬

If you’ve used nohuman before, I highly recommend updating to v0.5.0 and re-downloading the new DB.
Repo: github.com/mbhall88/nohuman
Keep your metagenomes clean 🧹🧬
So I rebuilt release 1 without masking, and added a release 2 database with no masking. The improvement in detection accuracy was substantial:

So I rebuilt release 1 without masking, and added a release 2 database with no masking. The improvement in detection accuracy was substantial:
Thanks for the great questions and discussion
Thanks for the great questions and discussion
There is likely inherent biases though based on error rates in reads for the kmer based methods
There is likely inherent biases though based on error rates in reads for the kmer based methods
-Thanks!
- See other thread where I have answered this
-Thanks!
- See other thread where I have answered this

(installable from wherever you get your podcasts 😉)

(installable from wherever you get your podcasts 😉)


LRGE is open-source and ready to integrate into your workflows as a Rust library or CLI application. Whether you’re on a high-performance cluster or a basic laptop, LRGE delivers fast and reliable genome size estimates. Get it here: github.com/mbhall88/lrge
LRGE is open-source and ready to integrate into your workflows as a Rust library or CLI application. Whether you’re on a high-performance cluster or a basic laptop, LRGE delivers fast and reliable genome size estimates. Get it here: github.com/mbhall88/lrge
LRGE uses significantly less CPU and memory than traditional approaches, making it ideal for both high-performance clusters and resource-limited environments.
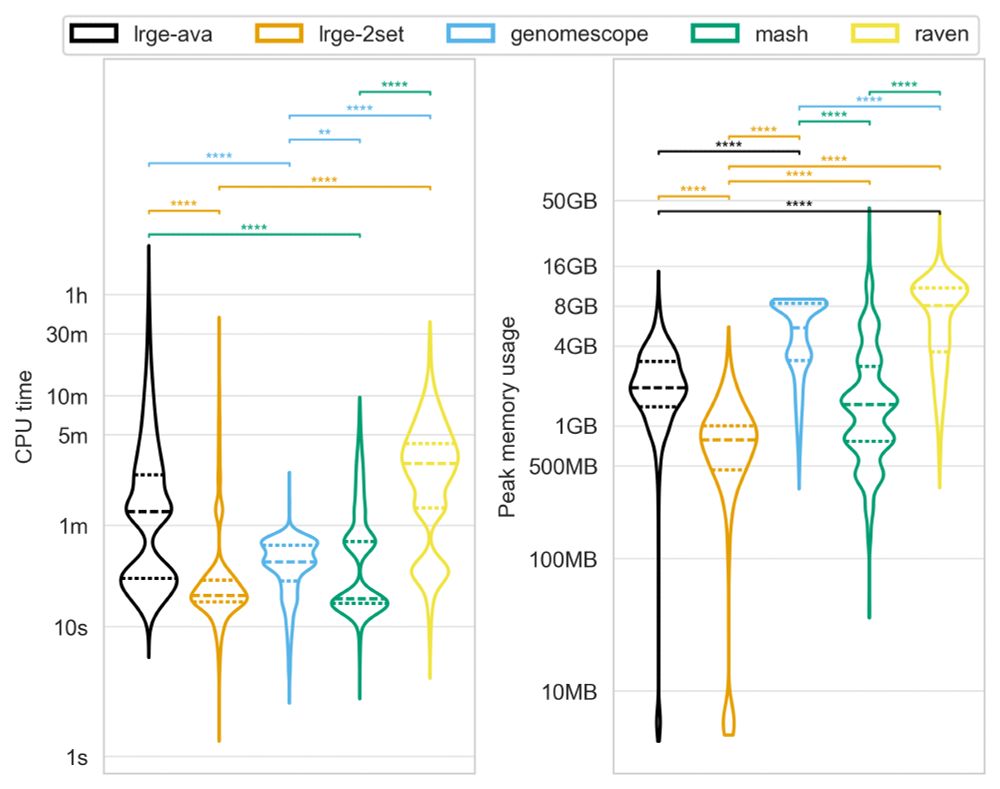
LRGE uses significantly less CPU and memory than traditional approaches, making it ideal for both high-performance clusters and resource-limited environments.
LRGE delivers estimates as reliable as assembly-based methods and better than k-mer-based approaches.
Relative error (y-axis) measures the proportional difference between the estimated and true genome size.
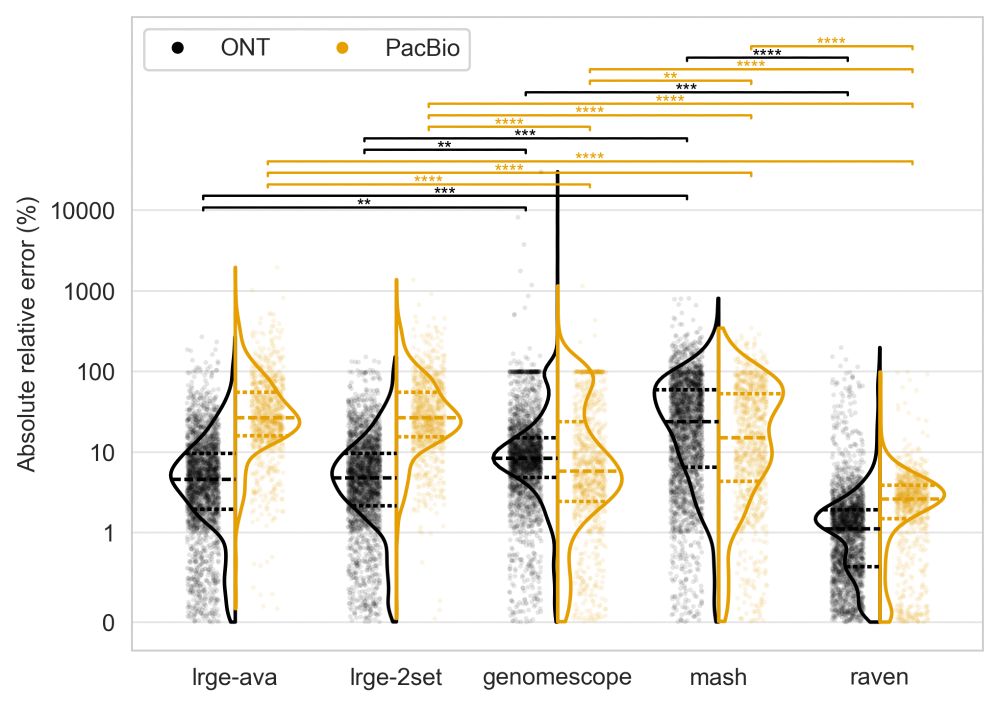
LRGE delivers estimates as reliable as assembly-based methods and better than k-mer-based approaches.
Relative error (y-axis) measures the proportional difference between the estimated and true genome size.

