Kir2.1: abundance → surface expression
PDZ3: abundance → CRIPT binding
KRAS: abundance → RAF1_RBD binding
Cosmos clearly separates direct binding residues from those with indirect effects.
[6/n]

Kir2.1: abundance → surface expression
PDZ3: abundance → CRIPT binding
KRAS: abundance → RAF1_RBD binding
Cosmos clearly separates direct binding residues from those with indirect effects.
[6/n]
Cosmos aggregates mutation effects by position, learns interpretable causal graphs via Bayesian model selection, and outputs residue-level direct vs indirect effects.
[5/n]
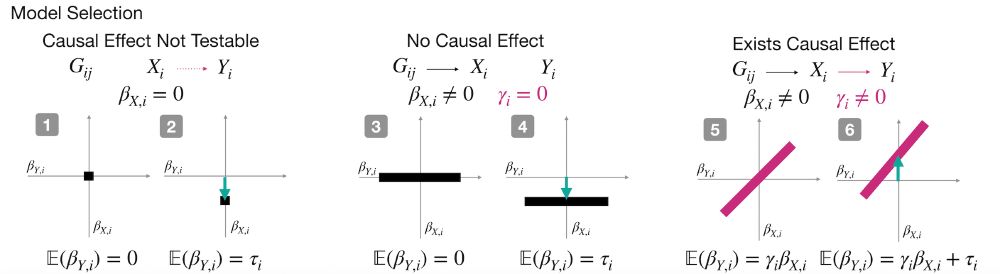
Cosmos aggregates mutation effects by position, learns interpretable causal graphs via Bayesian model selection, and outputs residue-level direct vs indirect effects.
[5/n]
1️⃣ Is there a causal link between the phenotypes?
2️⃣ How strong is it?
3️⃣ What would the downstream phenotype look like if we “normalized” the upstream one (i.e., counterfactual inference)?
[4/n]
1️⃣ Is there a causal link between the phenotypes?
2️⃣ How strong is it?
3️⃣ What would the downstream phenotype look like if we “normalized” the upstream one (i.e., counterfactual inference)?
[4/n]
No need for detailed biophysical models—just stats and data.
[3/n]

No need for detailed biophysical models—just stats and data.
[3/n]
But these phenotypes are often causally linked—e.g., when measuring activity, we may also capture effects propagated from abundance.
So how do we tell what’s direct vs indirect?
[2/n]
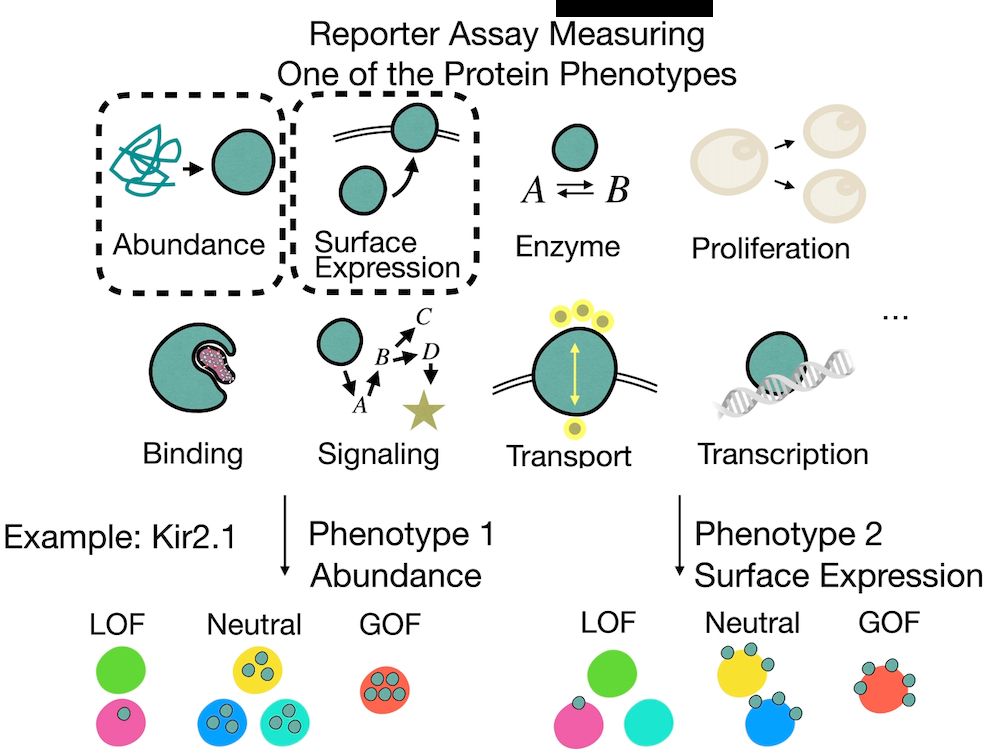
But these phenotypes are often causally linked—e.g., when measuring activity, we may also capture effects propagated from abundance.
So how do we tell what’s direct vs indirect?
[2/n]