
👏 Well deserved!
➡️ www.uni-wuerzburg.de/en/news-and-...
👏 Well deserved!
Thanks to everyone involved for this amazing collaborative effort to combine #proteomics with #functional #analysis.

Thanks to everyone involved for this amazing collaborative effort to combine #proteomics with #functional #analysis.
1️⃣ Appreciating the fundamental differences in scale and complexity between proteomics & transcriptomics
2️⃣ Recognizing the challenges without being intimidated. Being courageous to be creative.
blog.slavovlab.net/2023/09/23/t...

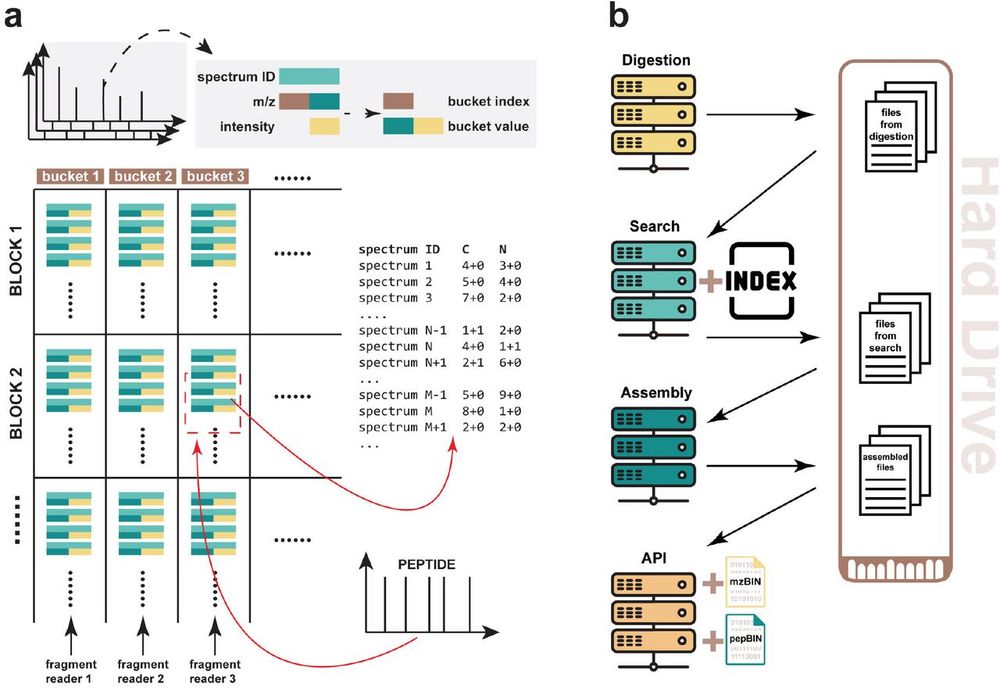
We go deeper into the #proteome, faster than ever!!
Exciting times ahead.. ☀️

It enabled many studies, but suitable tags have remained a limitation.
Soon, @parallelsq.bsky.social will introduce tags that improve multiplexing, sensitivity & coverage, helping realize huge potential pubs.acs.org/doi/10.1021/...
Stay tuned!
It enabled many studies, but suitable tags have remained a limitation.
Soon, @parallelsq.bsky.social will introduce tags that improve multiplexing, sensitivity & coverage, helping realize huge potential pubs.acs.org/doi/10.1021/...
Stay tuned!
📄 nature.com/articles/s41...
🌐 search.foldseek.com
📄 nature.com/articles/s41...
🌐 search.foldseek.com
"Brainwide silencing of prion protein by AAV-mediated delivery of an engineered compact epigenetic editor". #ShareGoodNewsToo

"Brainwide silencing of prion protein by AAV-mediated delivery of an engineered compact epigenetic editor". #ShareGoodNewsToo


www.nature.com/articles/s41...

www.nature.com/articles/s41...
I think we are slowly getting there. At least one step at a time, thanks to technological feats gained in MS community. For e.g recent nanoSPLITS method by @nanopots.bsky.social
The future is exciting!

www.nature.com/articles/s41...
I think we are slowly getting there. At least one step at a time, thanks to technological feats gained in MS community. For e.g recent nanoSPLITS method by @nanopots.bsky.social
The future is exciting!
www.biorxiv.org/content/10.1...

www.biorxiv.org/content/10.1...



I think you’ll love it :) @odedrechavi.bsky.social

I think you’ll love it :) @odedrechavi.bsky.social
It explores how normalisation, the CV formula and software parameters affect the outputs. It suggests parameters to use for biological and technical studies and provides an R package to calculate CVs.
doi.org/10.1021/acs....
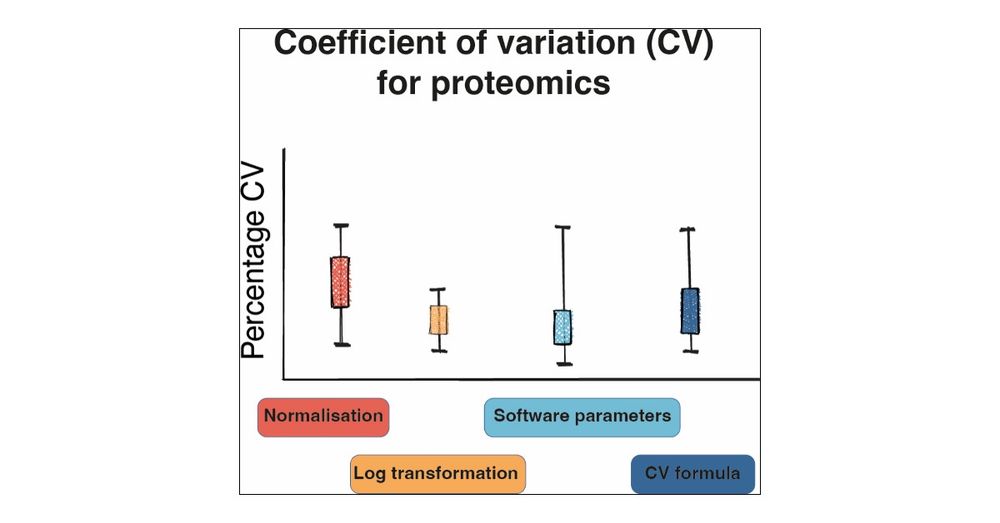
It explores how normalisation, the CV formula and software parameters affect the outputs. It suggests parameters to use for biological and technical studies and provides an R package to calculate CVs.
doi.org/10.1021/acs....

Check out this fantastic 15-minute intro video by @deeptijkundu.bsky.social, PRIDE's lead data curator:
📹 www.youtube.com/watch?v=VRNu...
#DataTransfer #Proteomics #Globus #Faster

Check out this fantastic 15-minute intro video by @deeptijkundu.bsky.social, PRIDE's lead data curator:
📹 www.youtube.com/watch?v=VRNu...
#DataTransfer #Proteomics #Globus #Faster
Here, they train a flow-matching model that can be conditioned by atomic coordinates and then use it to design enzymes based on active site geometry specified by density functional theory.
www.biorxiv.org/content/10.1...



Here, they train a flow-matching model that can be conditioned by atomic coordinates and then use it to design enzymes based on active site geometry specified by density functional theory.
www.biorxiv.org/content/10.1...
---
#proteomics #prot-other

---
#proteomics #prot-other
---
#proteomics #prot-preprint

---
#proteomics #prot-preprint

