1/9
www.nature.com/articles/s41...
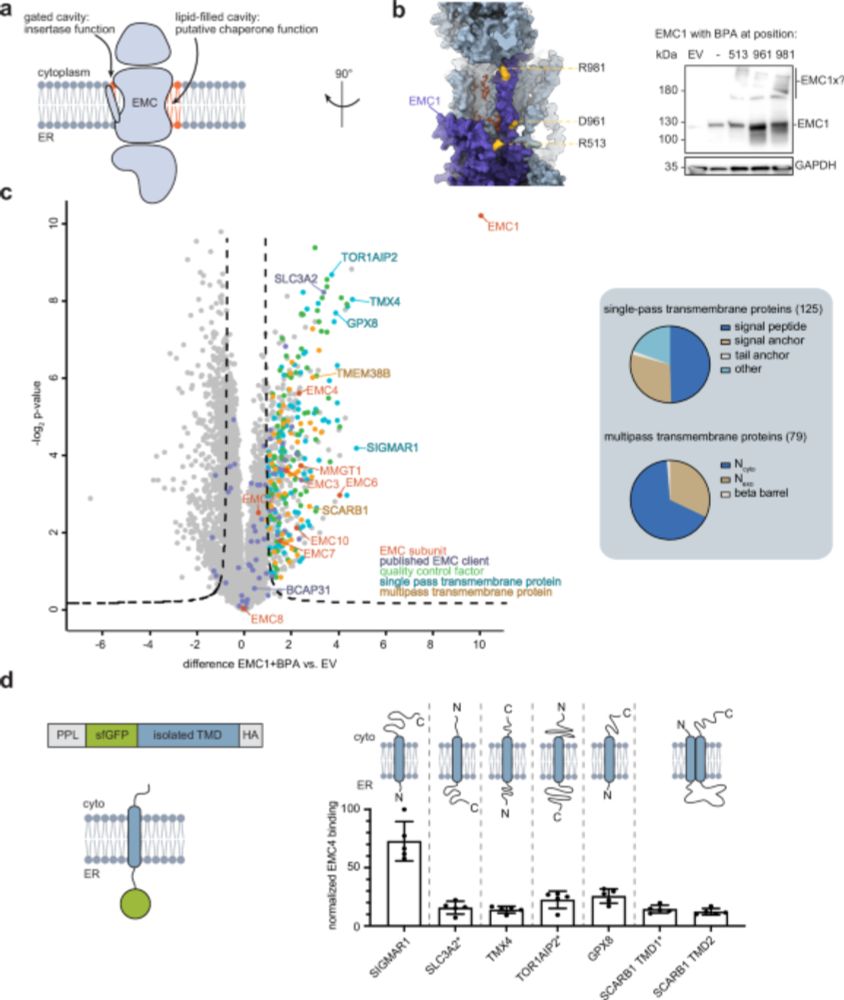
1/9
www.nature.com/articles/s41...

1/9
www.nature.com/articles/s41...
@natsmb.nature.com
1/7
www.nature.com/articles/s41...
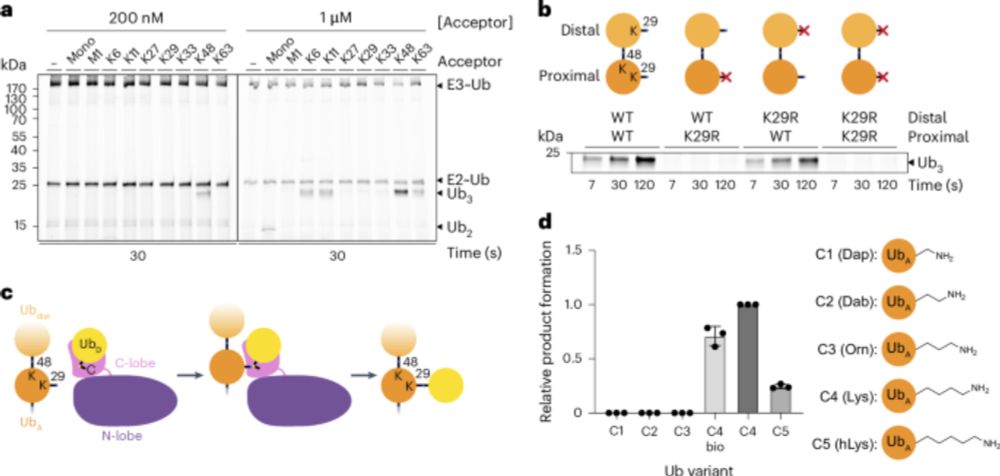
@natsmb.nature.com
1/7
www.nature.com/articles/s41...

@natsmb.nature.com
1/7
www.nature.com/articles/s41...
“𝗨𝗯𝗶𝗥𝗘𝗔𝗗 𝗱𝗲𝗰𝗶𝗽𝗵𝗲𝗿𝘀 𝗽𝗿𝗼𝘁𝗲𝗮𝘀𝗼𝗺𝗮𝗹 𝗱𝗲𝗴𝗿𝗮𝗱𝗮𝘁𝗶𝗼𝗻 𝗰𝗼𝗱𝗲 𝗼𝗳 𝗵𝗼𝗺𝗼𝘁𝘆𝗽𝗶𝗰 𝗮𝗻𝗱 𝗯𝗿𝗮𝗻𝗰𝗵𝗲𝗱 𝗞48 𝗮𝗻𝗱 𝗞63 𝘂𝗯𝗶𝗾𝘂𝗶𝘁𝗶𝗻 𝗰𝗵𝗮𝗶𝗻𝘀”
@cp-molcell.bsky.social
1/8
www.cell.com/molecular-ce...
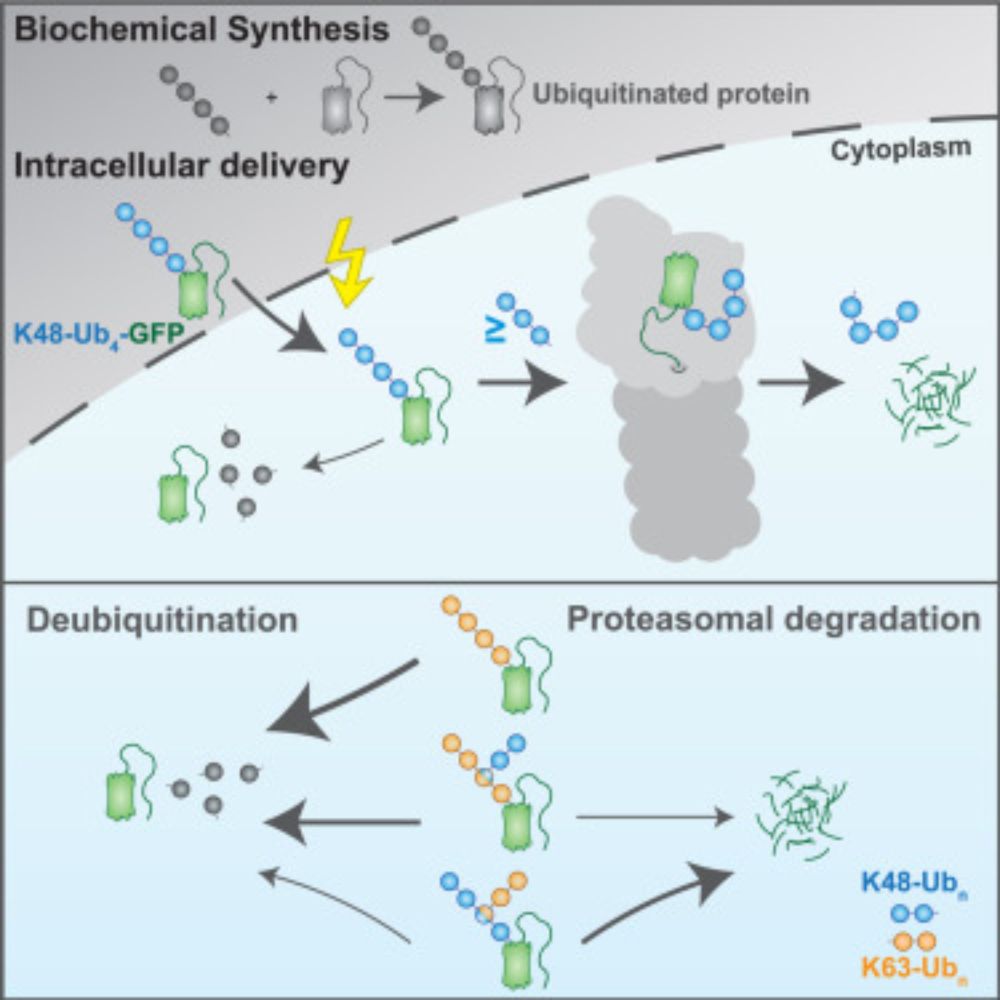
“𝗨𝗯𝗶𝗥𝗘𝗔𝗗 𝗱𝗲𝗰𝗶𝗽𝗵𝗲𝗿𝘀 𝗽𝗿𝗼𝘁𝗲𝗮𝘀𝗼𝗺𝗮𝗹 𝗱𝗲𝗴𝗿𝗮𝗱𝗮𝘁𝗶𝗼𝗻 𝗰𝗼𝗱𝗲 𝗼𝗳 𝗵𝗼𝗺𝗼𝘁𝘆𝗽𝗶𝗰 𝗮𝗻𝗱 𝗯𝗿𝗮𝗻𝗰𝗵𝗲𝗱 𝗞48 𝗮𝗻𝗱 𝗞63 𝘂𝗯𝗶𝗾𝘂𝗶𝘁𝗶𝗻 𝗰𝗵𝗮𝗶𝗻𝘀”
@cp-molcell.bsky.social
1/8
www.cell.com/molecular-ce...

